News

Research and Innovation |
HKUST Chemists Discover a “Super” Photoreductant to Leap Forward in Photocatalysis
Chemists at the School of Science of the Hong Kong University of Science and Technology (HKUST) have recently made a significant…
Research and Innovation |
Seven HKUST Projects Secure Funding in RAISe+ Second Round
The Hong Kong University of Science and Technology (HKUST) has shone in the second round of Research, Academic and Industry Sectors One-…

Research and Innovation |
HKUST Scientists Achieve Breakthrough in Light Manipulation Using Gyromagnetic Zero-Index Metamaterials
The Hong Kong University of Science and Technology (HKUST)-led research team has adopted gyromagnetic double-zero-index metamaterials (…
Research and Innovation |
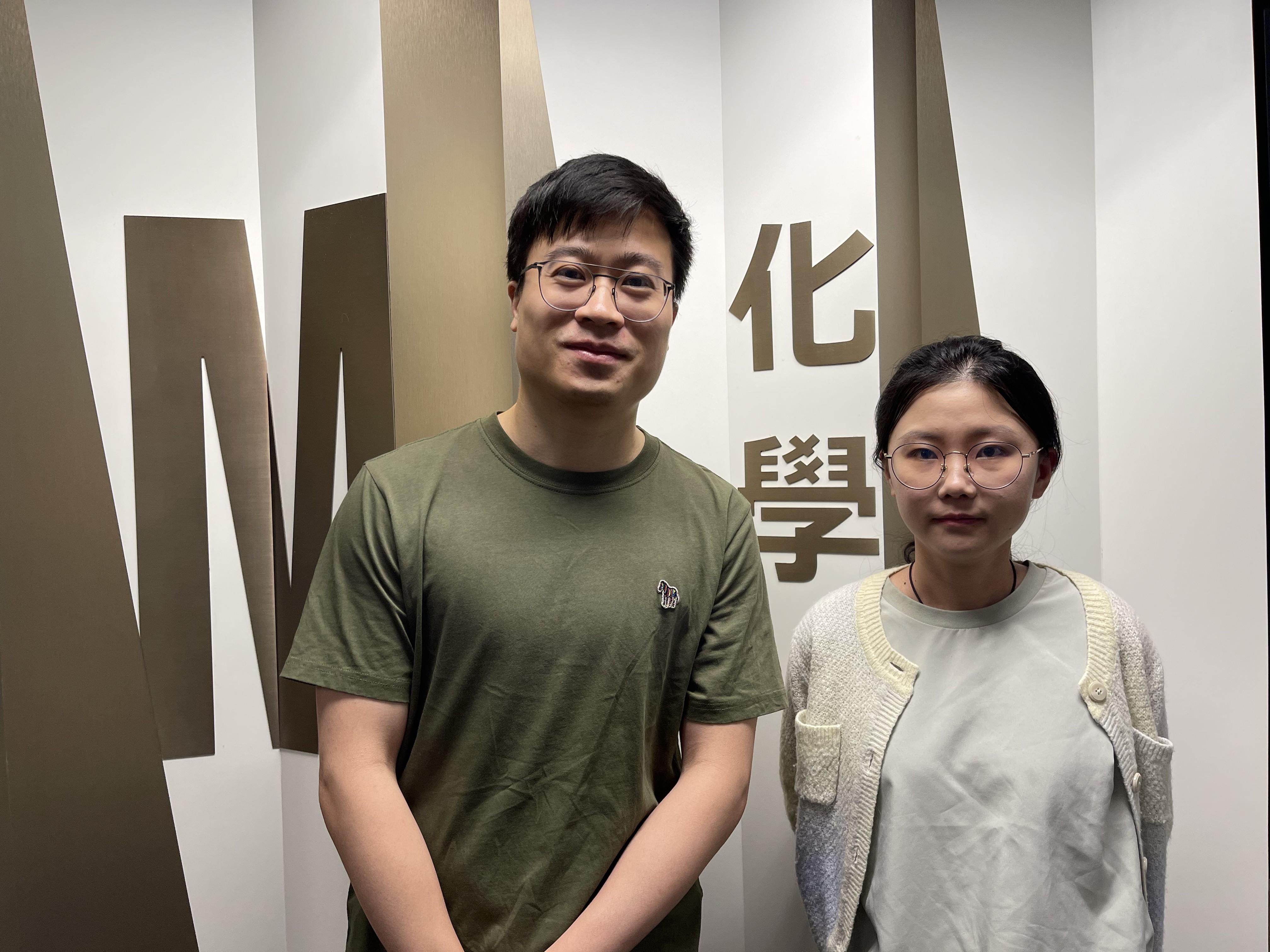
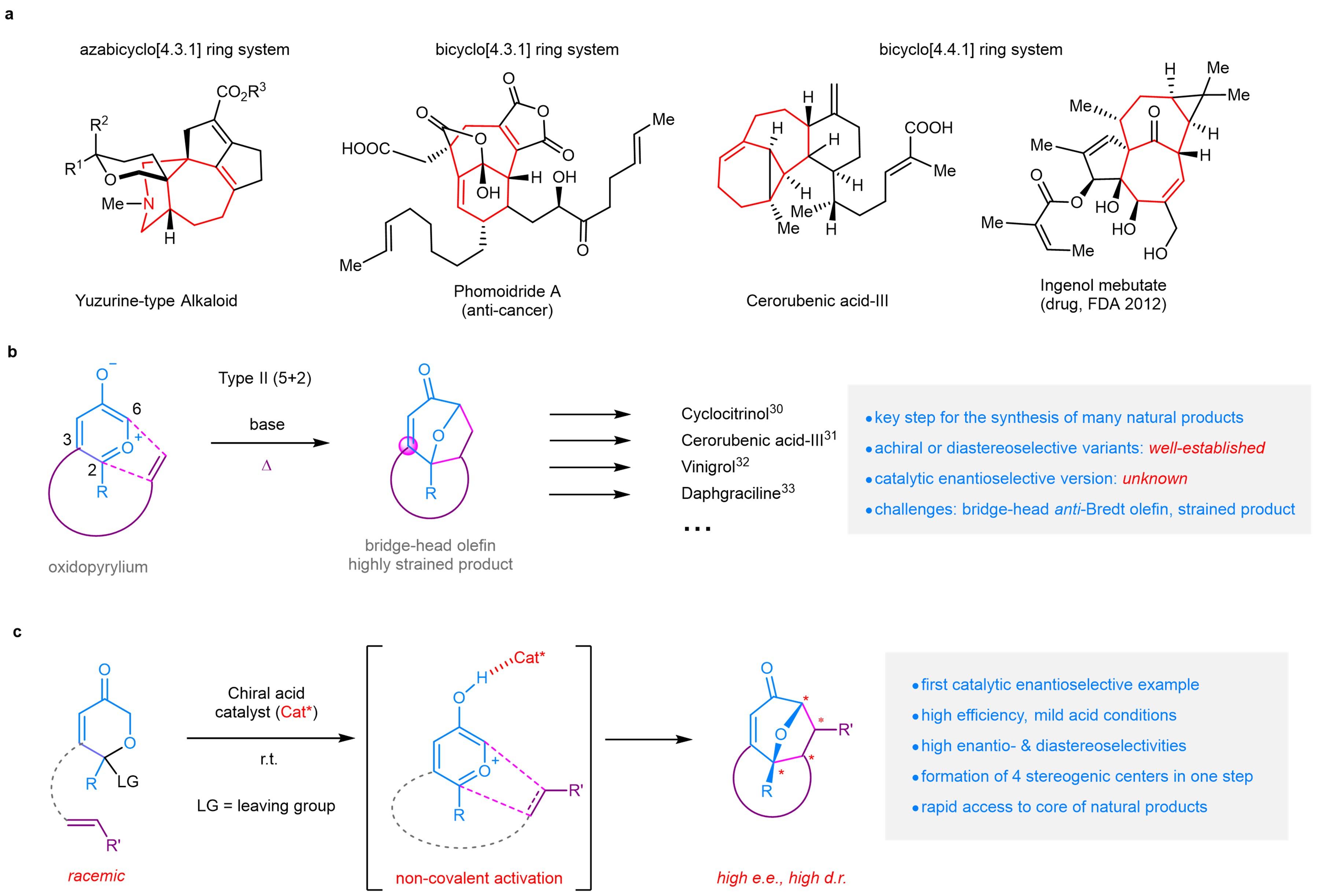
HKUST Chemists Innovate in the Synthesis of Chiral Bridged Polycyclic Compounds
(This article was originally published on EurekAlert! on June 4, 2025)
A Hong Kong University of Science and Technology (HKUST) research…

Recognition |
Recognition of Outstanding Faculty: Excellence in Research, Teaching, and Service
The School of Science Awards 2024/25 were presented to faculty members and teaching track faculty in recognition of their exceptional…
Research and Innovation |
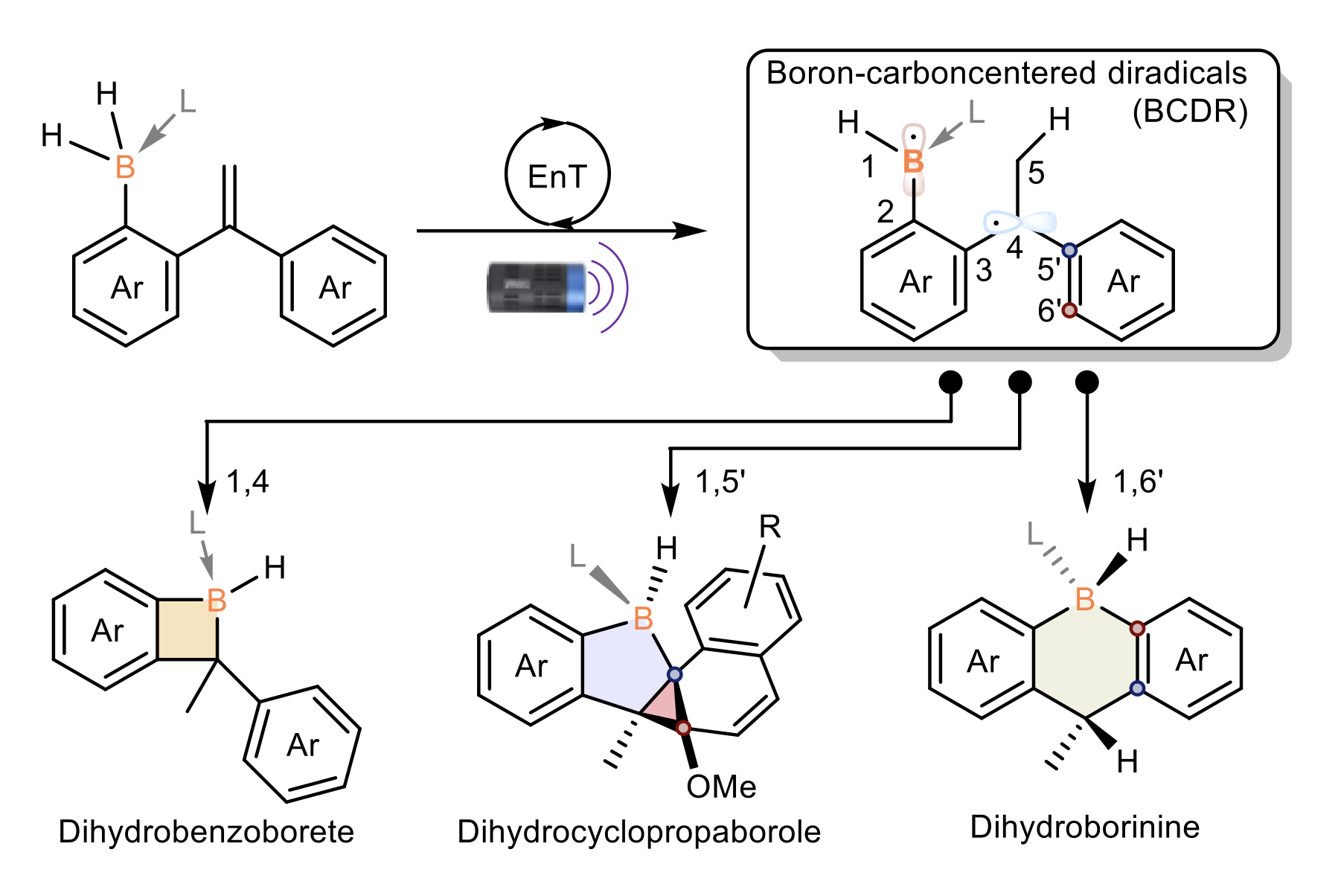
Unveiling Boracycles: HKUST Chemists Develop a New Approach to Synthesizing Strained Boracycles
(This article was originally published on EurekAlert! on May 19, 2025)
Boracycles are important functional scaffolds, finding increasing…

Recognition |
Quantum Leap: HKUST Physicist Awarded HKD 5 Million To Unlock Secrets of "Perfect" Materials and Revolutionize Energy Tech
Prof. SONG Xueyang, Assistant Professor of the Department of Physics at The Hong Kong University of Science and Technology (HKUST), has…
Research and Innovation |
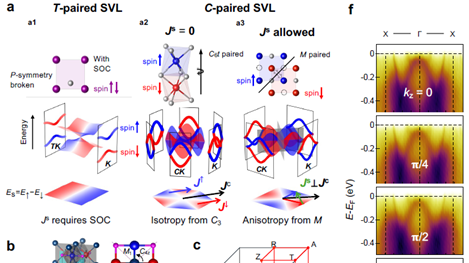
New Era of Magnetization: HKUST Research Sheds Light on Future Applications in Spintronics and Valleytronics
(This article was originally published on EurekAlert! on April 27, 2025)
Altermagnets, which exhibit momentum-dependent spin splitting…
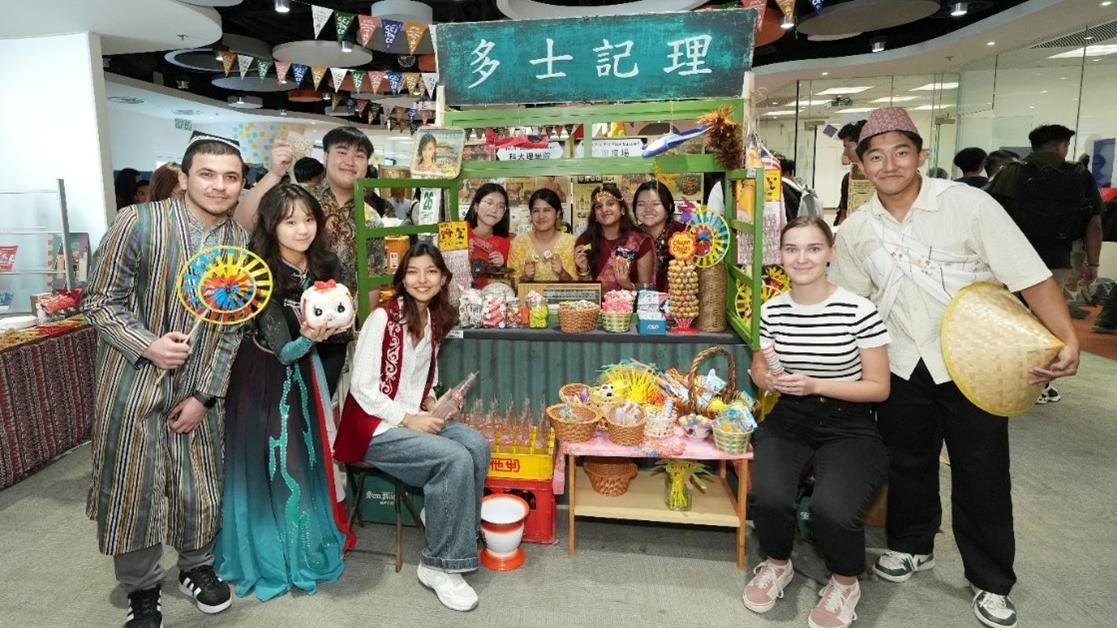

Student development & Bonding |
HKUST Community Unites for the School of Science's International Cultural Fair, Celebrating Global Diversity
Building on the vibrant experiences of previous years, the HKUST School of Science hosted the International Cultural Fair once again on…
Research and Innovation |
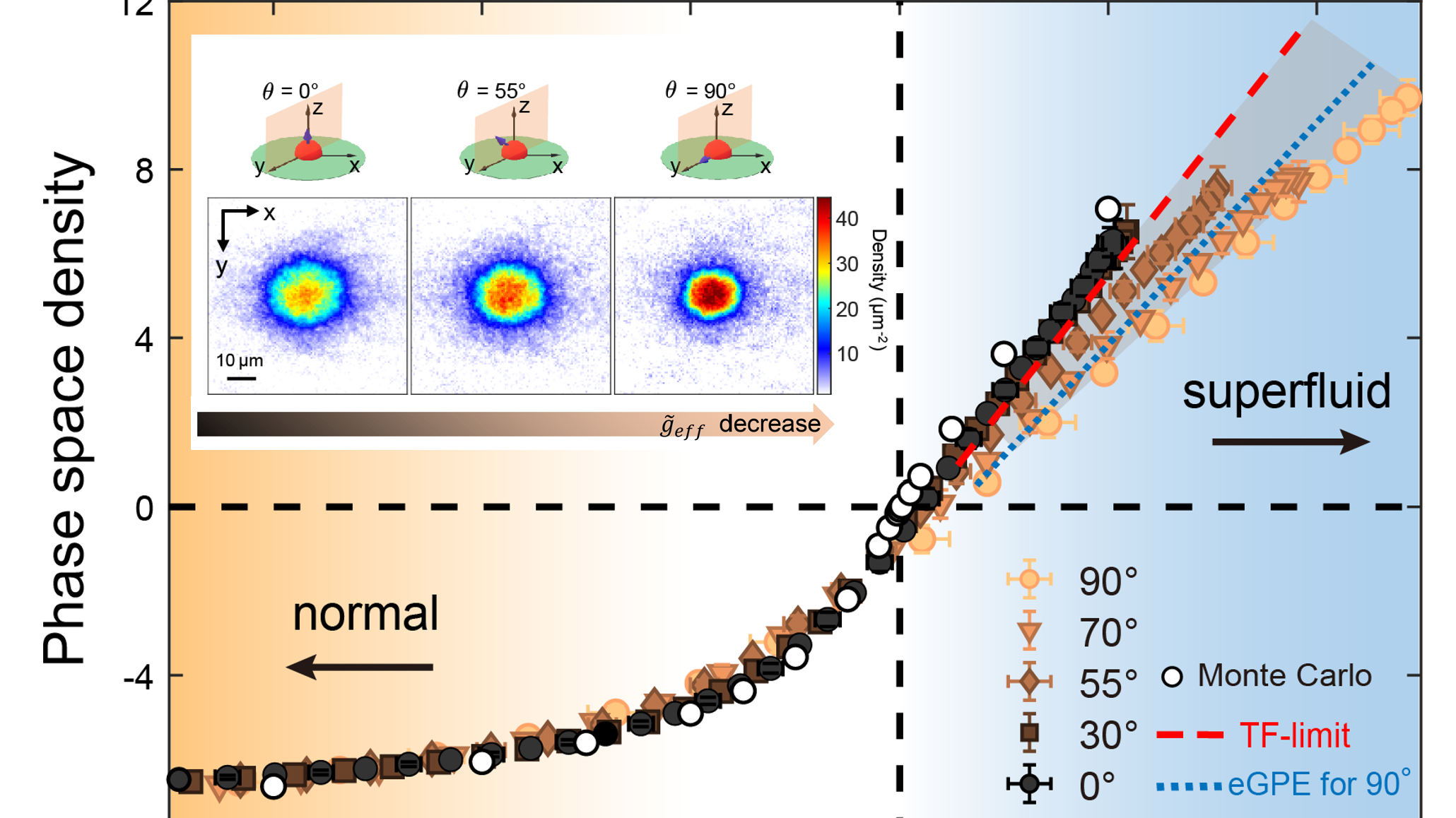
Unlocking the Secrets of Superfluid: HKUST Scientists Unveil How Dipolar Interactions Shape Two-Dimensional Superfluid Behavior
(This article was originally published on EurekAlert! on April 17, 2025)
In a recent study, an international team of physicists, led by …
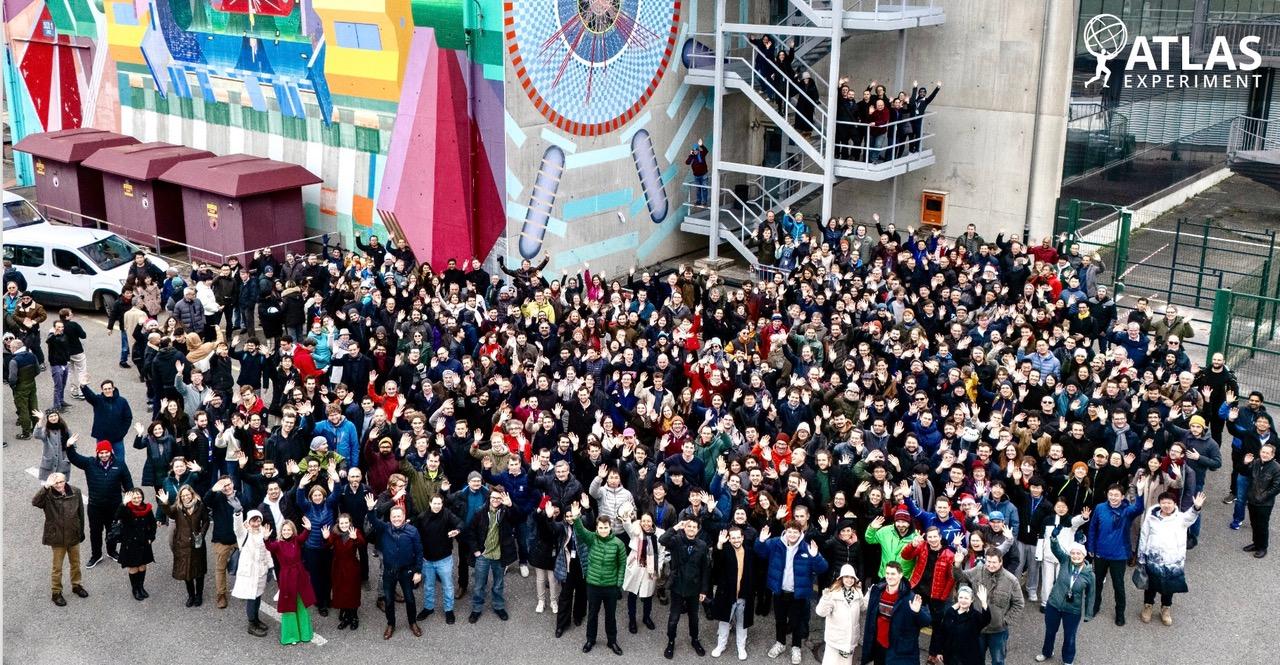

Recognition |
"Oscars of Science" Announces Winners: HKUST Celebrates Shared Victory in Breakthrough Prize in Fundamental Physics
Scientists from the Hong Kong University of Science and Technology (HKUST) are among the researchers worldwide honored with the 2025…

Research and Innovation |
HKUST Life Science Researchers Establish an Intracranial Optic Tract Injury Model to Reveal Mechanisms of Functional Circuit Reconstruction Following CNS Injury
(This article was originally published on EurekAlert! on March 28, 2025)
A research team led by Prof. LIU Kai from the Division of Life…